|
|
| Line 50: |
Line 50: |
| | *# Extract statistics from COMPARISON | | *# Extract statistics from COMPARISON |
| | | | |
| − | | [[File:GelDosimetrySlicelet_0.1_LandmarkRegistrationStep.png|800px|SlicerRT highlights]] | + | | [[File:GelDosimetryAnalysis_0.1.2_Step4Ui.png|800px|SlicerRT highlights]] |
| | |} | | |} |
| | | | |
Revision as of 18:54, 2 February 2014
Home < Documentation < Nightly < Modules < GelDosimetry



Introduction and Acknowledgements
Authors: Jennifer Andrea (PerkLab, Queen's University), Csaba Pinter (PerkLab, Queen's University), Mattea Welch (University of Toronto)
Contributors: Kevin Alexander (Kingston General Hospital), John Schreiner (Kingston General Hospital)
Contacts:
License: Slicer license
Download/install: install 3D Slicer, start 3D Slicer, open the Extension Manager, install the GelDosimetry extension
Extension Description
|

|
- Gel dosimetry analysis is a tool used in commissioning new radiation techniques and to validate the accuracy of radiation treatment by enabling visual comparison of the planned dose to the delivered dose, where correspondence between the two dose distributions is achieved using embedded landmarks. Gel dosimetry is based on imaging chemical systems spatially fixed in gelatin, which exhibit a detectable change upon irradiation. This chemical change is related to the amount of radiation received and can be probed by several imaging techniques, allowing 3D dose information to be obtained.
|
|
Use Cases
- Gel dosimetry analysis workflow:
- Loading planning image (PLANCT) and a planned dose distribution (PLANDOSE)
- Load an on-board imager cone beam CT scan (OBI)
- Register it to PLANCT
- Transform PLANDOSE using result transform (to OBI coordinate frame)
- Load 3D optical CT scan of the gel, or a 2D scan of the film. (MEASURED)
- Register MEASURED to OBI using fiducial marks, using rigid transform
- Transform MEASURED using result transform (to OBI coordinate frame)
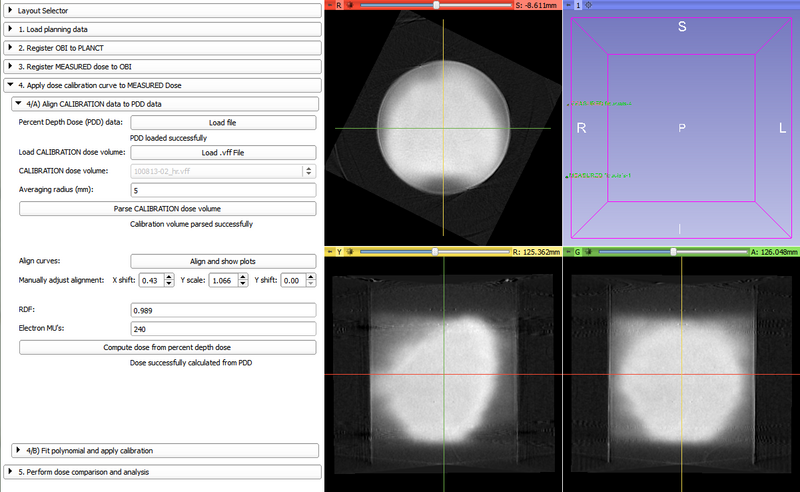
- Load Percent Depth Dose data (two-column spreadsheet)
- Load experimental CALIBRATION 3D optical CT scan from VFF file
- Get mean optical densities from the central cylinder of the CALIBRATION volume
- Align PDD curve with the curve resulting from step 10
- Calibrate PDD data using RDF and electron MU values input by the user
- Fit polynomial on the optical density vs. dose curve (resulting from the aligned calibration curves)
- Perform a gamma, chi, etc. test between MEASURED and PLANDOSE, yielding another 3D volume COMPARISON
- Extract statistics from COMPARISON
|
|
Tutorials
Similar Extensions
- SlicerRT: GelDosimetry uses several modules from the SlicerRT extension
References
Information for Developers




