Difference between revisions of "Slicer3.4:Training"
From Slicer Wiki
| Line 53: | Line 53: | ||
|- | |- | ||
| style="background:#FFD700; color:black" align="Center"| <span id="1.1"></span> '''Specialized''' | | style="background:#FFD700; color:black" align="Center"| <span id="1.1"></span> '''Specialized''' | ||
| − | | style="background:#FFFE59; color:black" | '''[[Media: | + | | style="background:#FFFE59; color:black" | '''[[Media:3DVisualization_SoniaPujol_RSNA2008.ppt | LiverSegmentation Tutorial]]''' <br><br> This tutorial introduces translational clinical scientists to the capabilities of the 3D Slicer software through the segmentation of the liver. |
'''Audience''': All users and developers. | '''Audience''': All users and developers. | ||
| style="background:#FFFE59; color:black" | '''Training data download is integrated with the LiverSegmentation tutorial (see tutorial slides)''' <br><br> | | style="background:#FFFE59; color:black" | '''Training data download is integrated with the LiverSegmentation tutorial (see tutorial slides)''' <br><br> | ||
Revision as of 22:33, 9 November 2009
Home < Slicer3.4:TrainingContents
Slicer 3.4 Tutorials
- The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment ( version 3.4) to accomplish certain tasks.
| Category | Tutorial | Sample Data | |
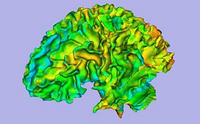
| Basic | Slicer3Minute Tutorial The Slicer3Minute tutorial is an introduction to the advanced 3D visualization capabilities of Slicer3.4. Audience: First time users. |
Slicer3Minute dataset The Slicer3Minute dataset contains an MR scan of the brain and 3D reconstructions of the anatomy |

|
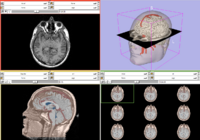
| Core | Slicer3Visualization Tutorial The Slicer3Visualization tutorial guides through 3D data loading and visualization in Slicer3.4. Audience: All beginning users including clinicians, scientists, engineers and programmers. |
Slicer3Visualization dataset The Slicer3VisualizationDataset contains two MR scans of the brain, a pre-computed labelmap and 3D reconstructions of the anatomy. |
|
| Core | Programming in Slicer3 Tutorial The Programming in Slicer3 tutorial is an introduction to the the integration of stand-alone programs outside of the Slicer3 source tree. Audience: Programmers and Engineers. |
HelloWorld Plugin The HelloWorld tutorial dataset contains an MR scan of the brain and pre-computed xml and C++ files for integrating the Hello World plug-in to Slicer3. |
|
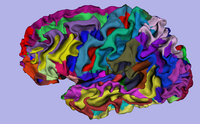
| Specialized | 3D Visualization of FreeSurfer Data The course guides through 3D visualization of FreeSurfer brain segmentations, surface reconstruction and parcellation results in Slicer3.4. Audience: All users. |
FreeSurfer Tutorial dataset The FreeSurfer dataset contains an MR scan of the brain and pre-computed FreeSurfer segmentation and cortical surface reconstructions. |
|
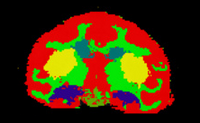
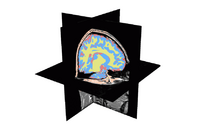
| Specialized | Automatic Segmentation Tutorial The course guides through the process of using the Expectation-Maximization Segmentation algorithm to automatically segment brain structures from MRI data. Audience: Programmers and Engineers. |
Automatic Segmentation dataset |
|
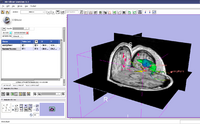
| Specialized | Neurosurgical Planning Tutorial This tutorial takes the trainee through a complete workup for neurosurgical patient-specific mapping. Also see this tutorial for information on how to use Slicer's affine registration, simple region growing, model maker and tractography modules. Audience: All users interested in image-guided therapy. |
Neurosurgical Planning dataset |
|
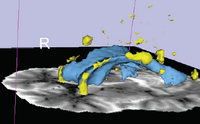
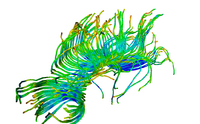
| Specialized | Diffusion MRI Tutorial This tutorial guides you through the process of loading diffusion MR data, estimating diffusion tensors, and performing tractography of white matter bundles. Audience: All users and developers. |
Diffusion dataset |

|
| Specialized | ChangeTracker Tutorial This tutorial describes the use of ChangeTracker module to detect changes in tumor volume from two MRI scans. Audience: All users interested in longitudinal analysis of pathology. |
Training data download is integrated with the ChangeTracker module (see tutorial slides) |
|
| Specialized | LiverSegmentation Tutorial This tutorial introduces translational clinical scientists to the capabilities of the 3D Slicer software through the segmentation of the liver. Audience: All users and developers. |
Training data download is integrated with the LiverSegmentation tutorial (see tutorial slides) |
Slicer Tutorial Contest
(under construction)
The following tutorials were part of the Summer 2009 Slicer tutorial contest and Winter 2009 Slicer tutorial contest.
Software Installation
- The Slicer download page contains information on how to obtain a compiled version of Slicer for a variety of platforms and where to find the source code for Slicer 3.
Software Documentation
- For the Slicer 3.4 manual pages please click here. These pages are the reference manual for Slicer 3.4 and briefly explain the functionality found in panels and modules.
Older Tutorials
- Visit the Slicer 3.2 training page.
- Visit the Slicer 2 training page.