Difference between revisions of "Modules:StochasticTractography-Documentation-3.4"
| Line 105: | Line 105: | ||
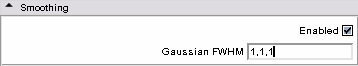
* '''Smoothing panel:''' | * '''Smoothing panel:''' | ||
** Gaussian FWHM: this filter defines a Full Width Half Maximum. You can define it for each direction in modifying each component of the 3-vector. | ** Gaussian FWHM: this filter defines a Full Width Half Maximum. You can define it for each direction in modifying each component of the 3-vector. | ||
| − | ** Advice: you can enable solely that functionality and compute several times with different parameters till | + | ** Advice: you can enable solely that functionality and compute several times with different parameters till satisfaction. |
{| | {| | ||
|[[Image:Smoothmenu.png|thumb|500px|Smoothing step]] | |[[Image:Smoothmenu.png|thumb|500px|Smoothing step]] | ||
| Line 114: | Line 114: | ||
| | | | ||
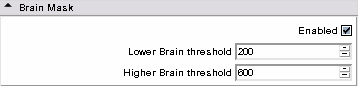
* '''Brain Mask panel:''' | * '''Brain Mask panel:''' | ||
| − | ** Lower/Higher Brain Threshold: this filter is based on a simple level set segmentation - intensities of the baseline lying between the two values will be represented, others are set to 0. The most common use of this filter is to remove ventricles from the tractography domain and most of the outside of the brain. | + | ** Lower/Higher Brain Threshold: this filter is based on a simple level set segmentation - intensities of the baseline lying between the two values will be represented, others are set to 0. The most common use of this filter is to remove ventricles from the tractography domain and most of the outside of the brain. |
| + | ** Advice: you can enable solely that functionality and compute several times with different parameters till satisfaction. | ||
{| | {| | ||
|[[Image:Maskmenu.png|thumb|500px|Brain mask step]] | |[[Image:Maskmenu.png|thumb|500px|Brain mask step]] | ||
| Line 125: | Line 126: | ||
** Important: this step is not mandatory. It is here to evaluate the whole tensor and achieve measurements like FA, mode or trace. You did not need it to achieve tractography. | ** Important: this step is not mandatory. It is here to evaluate the whole tensor and achieve measurements like FA, mode or trace. You did not need it to achieve tractography. | ||
** FA, mode and trace are sent as scalar volumes and inserted in the MRML tree to be further used. | ** FA, mode and trace are sent as scalar volumes and inserted in the MRML tree to be further used. | ||
| + | ** Advice: you can enable solely that functionality and compute several times with different parameters till satisfaction. | ||
{| | {| | ||
|[[Image:Tensormenu.png|thumb|500px|Tensor step]] | |[[Image:Tensormenu.png|thumb|500px|Tensor step]] | ||
Revision as of 01:08, 16 April 2009
Home < Modules:StochasticTractography-Documentation-3.4Return to Slicer 3.4 Documentation
Module Name
Stochastic Tractography
General Information
Module Type & Category
Type: Interactive
Category: DTI
Authors, Collaborators & Contact
- Author: Julien von Siebenthal
- Contributor: Steve Pieper
- Contact: jvs@bwh.harvard.edu
Module Description
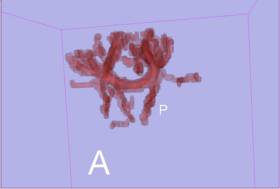
As a main purpose, the stochastic tractography module helps to evaluate connectivity between two regions of interest (ROIs) of the brain. These ROIs define generally grey matter regions having a specific neurophysiological function. Extensively, study involving more than two regions could still be done by pairing the regions two by two.
Usage
- You want to study fiber path from a single region of interest (ROI)
- You want to evaluate connectivity between two ROIs
Description
With the stochastic tractography module, you can:
|
|
|
|
|
|
Quick Tour of Features and Use
|
|
|
|
|
|
Development
Dependencies
Volumes
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
More Information
Acknowledgment
National Alliance for Medical Image Computing (NAMIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149 (to Ron Kikinis, Marek Kubicki).
References
- Björnemo M, Brun A, Kikinis R, Westin CF. Regularized stochastic white matter tractography using diffusion tensor MRI. In Fifth International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI'02). Tokyo, Japan, 2002;435-442.
- Friman, O., Farneback, G., Westin CF. A Bayesian Approach for Stochastic White Matter Tractography. IEEE Transactions on Medical Imaging, Vol 25, No. 8, Aug. 2006
- Shenton, M.E., Ngo, T., Rosenberger, G., Westin, C.F., Levitt, J.J., McCarley, R.W., Kubicki, M. Study of Thalamo-Cortical White Matter Fiber Tract Projections in Schizophrenia Using Diffusion Stochastic Tractography. Poster presented at the 46th Meeting of the American College of Neuropsychopharmacology, Boca Raton, FL, December 2007.