Difference between revisions of "Documentation/Nightly/Extensions/TCIABrowser"
| Line 29: | Line 29: | ||
[[Image:TCIABrowser_logo.png]] | [[Image:TCIABrowser_logo.png]] | ||
| − | The Cancer Imaging Archive (TCIA) hosts a large collection of Cancer medical imaging data which is available to public through a programmatic interface (REST API). TCIA Browser is a Slicer module by which the user can connect to the TCIA archive, browse different collections, | + | The Cancer Imaging Archive (TCIA) hosts a large collection of Cancer medical imaging data which is available to public through a programmatic interface (REST API). TCIA Browser is a Slicer module by which the user can connect to the TCIA archive, browse different collections, patient subjects, studies and series, download the images and visualize them in 3D Slicer. |
As of February 2014, the following collections of TCIA are available via public TCIA API interface (see [https://wiki.cancerimagingarchive.net/display/Public/Collections description of TCIA collections] for details): | As of February 2014, the following collections of TCIA are available via public TCIA API interface (see [https://wiki.cancerimagingarchive.net/display/Public/Collections description of TCIA collections] for details): | ||
| Line 72: | Line 72: | ||
{| | {| | ||
| − | * ''' | + | * '''Browsing Collections, patients and studies''' |
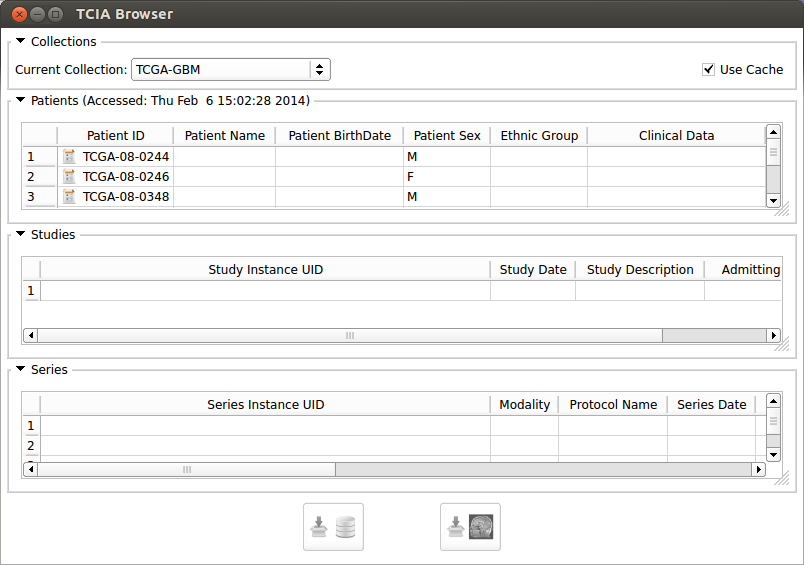
| − | + | By opening TCIABrowser module it will automatically connect to the the TCIA server and list all of the available collections. | |
| − | |||
|[[Image:TCIA-New01.png|thumb|360px|Module GUI]] | |[[Image:TCIA-New01.png|thumb|360px|Module GUI]] | ||
| − | |||
| − | |||
| − | |||
In the browser window from the combo box select a collection. The browser will get the patient data from the TCIA server and will populate the Patients table. In patient table, the clinical data icon besides the patient ID [[Image:Report-icon-tcia.png|thumb|50px|Clinical Data Icon]] means that the clinical data for that specific patient is available in TCGA. The use Cache check box will cache the query results on your hard drive which makes the next selections much faster. If you want to do the query (connect to TCIA Server) you can uncheck this box. Above the Patients table you can see the last access time. The process is the same for studies and series tables, selecting a patient will populate the study table for the selected one and selecting a study will update the series table. | In the browser window from the combo box select a collection. The browser will get the patient data from the TCIA server and will populate the Patients table. In patient table, the clinical data icon besides the patient ID [[Image:Report-icon-tcia.png|thumb|50px|Clinical Data Icon]] means that the clinical data for that specific patient is available in TCGA. The use Cache check box will cache the query results on your hard drive which makes the next selections much faster. If you want to do the query (connect to TCIA Server) you can uncheck this box. Above the Patients table you can see the last access time. The process is the same for studies and series tables, selecting a patient will populate the study table for the selected one and selecting a study will update the series table. | ||
| Line 87: | Line 83: | ||
{| | {| | ||
| − | * ''' | + | * '''Downloading Series''' |
| − | + | In the final step when you select a series the download icons will become activated. Pressing the "Download and Index" button will download the images from TCIA to your computer and index the DICOM files inside 3D Slicer DICOM database. So you can review them, check the meta-data and load into the scene later with Slicer DICOM module. Pressing the "Download and Load" button will download the images and load them into the Slicer scene. You can select multiple items from all of the tables. By holding the Ctrl key and clicking on different patients, the studies for all the selected ones will be added to the studies table. You can select all the studies by pressing 'Select All' button or make a specific selection by Ctrl+Click and all the available series for download will be added to the series table. At the final step you will select series for download from the series table. The total number of images for the selected series is indicated at the bottom right corner of series table. After pressing the download button you can check the download status of each series at the 'Download Status' collapsible button at the module's widget. While the download is in progress you can still browse and add other series to the download queue or view the downloaded images in the 3D Slicer Scene. | |
| − | + | [[Image:DownloadAndIndex.png|thumb|100px|Download and Index Button]][[Image:DownloadAndLoad.png|thumb|100px|Download and Load Button]] | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | In the final step when you select a series the download icons will become activated. Pressing the "Download and Index" button will download the images from TCIA to your computer and index the DICOM files inside 3D Slicer DICOM database. So you can review them, check the meta-data and load into the scene later with Slicer DICOM module. Pressing the "Download and Load" button will download the images and load them into the Slicer scene.[[Image:DownloadAndIndex.png|thumb|100px|Download and Index Button]][[Image:DownloadAndLoad.png|thumb|100px|Download and Load Button]] | ||
|[[Image:TCIA-New05.png|thumb|800px|Module GUI]] | |[[Image:TCIA-New05.png|thumb|800px|Module GUI]] | ||
| | | | ||
| + | |||
| + | |[[Image:Tcia-multi.png|thumb|800px|Multiple Selections]] | ||
| + | |[[Image:Tcia-download-status.png|thumb|400px|Download Status]] | ||
|} | |} | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| + | {| | ||
| + | * '''Access Clinical Data from TCGA''' | ||
| + | For some of the collections clinical data is available through The Cancer Genome Atlas (TCGA) website. The patients with this information are marked with a unique icon besides their patient ID. The clinical data can be accessed by right clicking on the Patient ID area to open the context menu and clicking "Get clinical data". | ||
| + | |[[Image:TCIA-New03.png|thumb|600px|Module GUI]] | ||
| + | |[[Image:TCIA-New04.png|thumb|600px|Module GUI]] | ||
|} | |} | ||
| + | {| | ||
| + | |||
| + | * '''Settings ''' | ||
| + | If you have your own unique API and want to use that you can add your own API using the "+" button and then select it using the combo box. | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Similar Modules}} | {{documentation/{{documentation/version}}/module-section|Similar Modules}} | ||
* [[Documentation/4.3/Extensions/XNATSlicer|XNATSlicer]] | * [[Documentation/4.3/Extensions/XNATSlicer|XNATSlicer]] | ||
*[[Documentation/Nightly/Modules/DICOM| DICOM]] | *[[Documentation/Nightly/Modules/DICOM| DICOM]] | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|References}} | {{documentation/{{documentation/version}}/module-section|References}} | ||
Revision as of 13:06, 18 May 2015
Home < Documentation < Nightly < Extensions < TCIABrowser
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
|
Extension: TCIABrowser License: Slicer License
|
Module Description
The Cancer Imaging Archive (TCIA) hosts a large collection of Cancer medical imaging data which is available to public through a programmatic interface (REST API). TCIA Browser is a Slicer module by which the user can connect to the TCIA archive, browse different collections, patient subjects, studies and series, download the images and visualize them in 3D Slicer.
As of February 2014, the following collections of TCIA are available via public TCIA API interface (see description of TCIA collections for details):
- QIBA CT-1C
- TCGA-GBM *
- RIDER PHANTOM PET-CT
- RIDER PHANTOM MRI
- Phantom FDA
- TCGA-LGG *
- TCGA-KIRC *
- TCGA-BRCA *
- BREAST-DIAGNOSIS
- PROSTATE-DIAGNOSIS
- LIDC-IDRI
- TCGA-LUAD *
- RIDER Lung CT
- RIDER NEURO MRI
- RIDER Lung PET-CT
- CT COLONOGRAPHY
- REMBRANDT
- RIDER Breast MRI
For the collections marked with *, available clinical data is linked from The Cancer Genome Atlas.
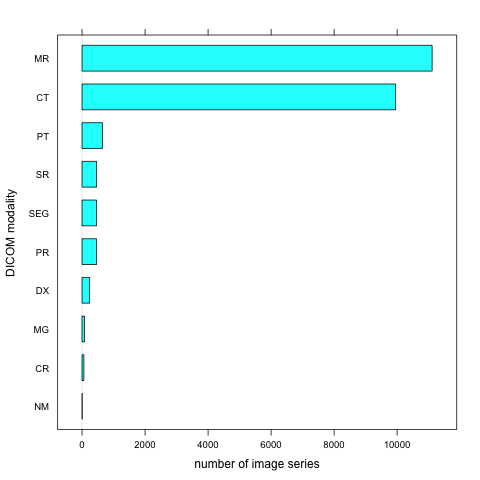
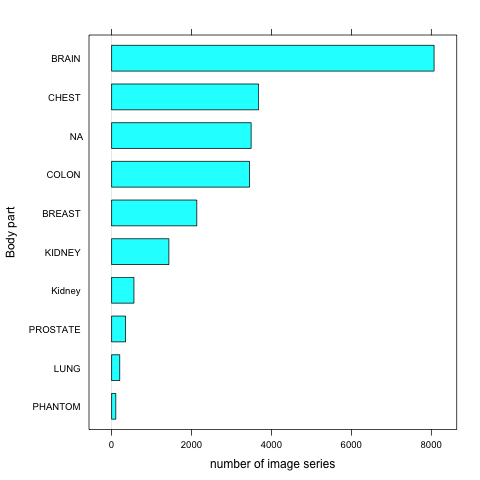
TCIABrowser gives you access to 18 collections, which contain a total of over 23,000 image series from a variety of manufacturers and of different modalities and organs. Some of the summary statistics of the data available from the public collections listed is provided below.
Tutorials
Not available at this time.
Panels and their use
- Browsing Collections, patients and studies
| In the browser window from the combo box select a collection. The browser will get the patient data from the TCIA server and will populate the Patients table. In patient table, the clinical data icon besides the patient ID means that the clinical data for that specific patient is available in TCGA. The use Cache check box will cache the query results on your hard drive which makes the next selections much faster. If you want to do the query (connect to TCIA Server) you can uncheck this box. Above the Patients table you can see the last access time. The process is the same for studies and series tables, selecting a patient will populate the study table for the selected one and selecting a study will update the series table. |
- Downloading Series
- Access Clinical Data from TCGA
- Settings
Similar Modules
References
- Quantitative Image Informatics for Cancer Research (QIICR)
- Quantitative Imaging Network (QIN)
- TCIA Home Page
- cBioPortal for Cancer Genomics Web Interface
- Description of TCIA Collections
- TCIA Rest API Documentation
- Project page at NAMIC 2014 Project Week
- Mashape page for testing TCIA API endpoint
Information for Developers
Tip: The REST API can be checked by REST Console Google Chrome extension.
| Section under construction. |