Difference between revisions of "Documentation/Nightly/Modules/SobolevSegmenter"
| Line 30: | Line 30: | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Use Cases}} | {{documentation/{{documentation/version}}/module-section|Use Cases}} | ||
| − | The Sobolev segmenter is general and can be used with any 2D data, as explained in the tutorial. | + | The Sobolev segmenter is a general image segmenter, and it can be used with any 2D data, as explained in the tutorial. |
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Tutorials}} | {{documentation/{{documentation/version}}/module-section|Tutorials}} | ||
# Load the image (input volume): [[File:DICOM_example.jpg|200 px]] | # Load the image (input volume): [[File:DICOM_example.jpg|200 px]] | ||
| − | # Use built in editor to select | + | # Use built in editor to select a single initial mask (or load a binary mask file): [[File:mask_example.jpg|200 px]] |
# Select Segmenation->SobolevSegmenter module | # Select Segmenation->SobolevSegmenter module | ||
# Choose the Input Volume and the Initial Mask accordingly. Create a new volume for the Output Volume. | # Choose the Input Volume and the Initial Mask accordingly. Create a new volume for the Output Volume. | ||
Revision as of 03:16, 25 February 2013
Home < Documentation < Nightly < Modules < SobolevSegmenterIntroduction and Acknowledgements
|
This work is part of the National Alliance for Medical Image Computing (NA-MIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research. Information on NA-MIC can be obtained from the NA-MIC website. | |||
|
Module Description
This extension implements Sobolev inner product based active contour, using Chan-Vese energy functional. The segmentation is appropriate for 2D images. The obtained parametric contour is generally smooth, but able to catch concavities.
Use Cases
The Sobolev segmenter is a general image segmenter, and it can be used with any 2D data, as explained in the tutorial.
Tutorials
- Load the image (input volume):
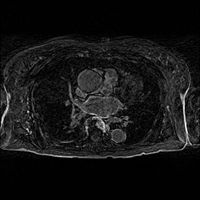
- Use built in editor to select a single initial mask (or load a binary mask file):
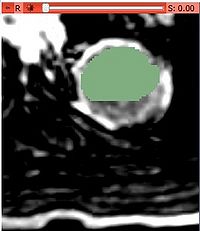
- Select Segmenation->SobolevSegmenter module
- Choose the Input Volume and the Initial Mask accordingly. Create a new volume for the Output Volume.
- Press Apply button.
- After a few second the following output volume should appear:
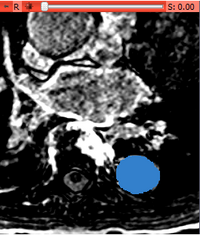
Panels and their use
The module has the following panel:
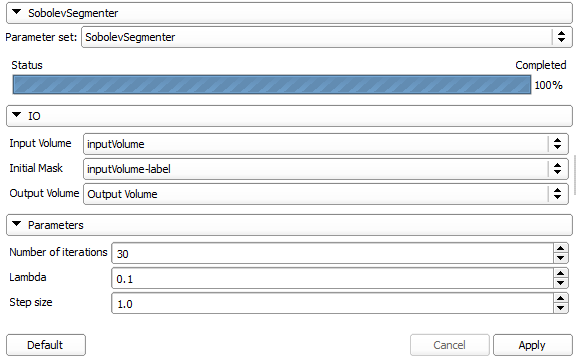 The IO section of this panel defines two input images (data and initial mask) and one output image (final mask). The algorithm has three parameters: self-explanatory number of iterations and contour evolution step size. In addition, the parameter lambda chooses the smoothness of the contour (smoothing kernel width).
The IO section of this panel defines two input images (data and initial mask) and one output image (final mask). The algorithm has three parameters: self-explanatory number of iterations and contour evolution step size. In addition, the parameter lambda chooses the smoothness of the contour (smoothing kernel width).
Similar Modules
N/A
References
N/A
Information for Developers
| Section under construction. |
