Difference between revisions of "Documentation/Nightly/Extensions/SlicerRT"
m (Text replacement - "\[http:\/\/www\.slicer\.org\/slicerWiki\/index\.php\/([^ ]+) ([^]]+)]" to "$2") |
|||
| (4 intermediate revisions by one other user not shown) | |||
| Line 56: | Line 56: | ||
**[[Documentation/{{documentation/version}}/Modules/SegmentMorphology|Segment morphology]] (Add/remove margin, Unify, Intersect, etc.) | **[[Documentation/{{documentation/version}}/Modules/SegmentMorphology|Segment morphology]] (Add/remove margin, Unify, Intersect, etc.) | ||
* I/O | * I/O | ||
| − | **[[Documentation/{{documentation/version}}/Modules/DicomRtImport|DICOM-RT import]], [ | + | **[[Documentation/{{documentation/version}}/Modules/DicomRtImport|DICOM-RT import]], [https://www.slicer.org/wiki/Documentation/Nightly/Modules/DICOM#DICOM_export export] (handles datasets of types RT Structure Set, RT Dose, RT Plan, RT Image) |
| + | **[[Documentation/{{documentation/version}}/Modules/DicomSroImport|DICOM-SRO import/export]] (handles DICOM Spatial Registration object, both rigid and deformable) | ||
*[https://github.com/SlicerRt/SlicerRT/tree/master/BatchProcessing Batch processing scripts] (currently only one is available for command-line conversion of RTSS to volume nodes) | *[https://github.com/SlicerRt/SlicerRT/tree/master/BatchProcessing Batch processing scripts] (currently only one is available for command-line conversion of RTSS to volume nodes) | ||
| Line 69: | Line 70: | ||
**[[Documentation/{{documentation/version}}/Modules/Data|Subject hierarchy]] | **[[Documentation/{{documentation/version}}/Modules/Data|Subject hierarchy]] | ||
**[[Documentation/{{documentation/version}}/Modules/Transforms|Transform visualizer]] | **[[Documentation/{{documentation/version}}/Modules/Transforms|Transform visualizer]] | ||
| − | **DICOM-RT export, as [ | + | **DICOM-RT export, as [[Documentation/Labs/DICOMExport|improved DICOM export function]] |
**[[Documentation/{{documentation/version}}/Modules/Segmentations|Segmentations]] | **[[Documentation/{{documentation/version}}/Modules/Segmentations|Segmentations]] | ||
**[[Documentation/{{documentation/version}}/Modules/SegmentEditor|Segment Editor]] | **[[Documentation/{{documentation/version}}/Modules/SegmentEditor|Segment Editor]] | ||
| Line 98: | Line 99: | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/extension-section|Tutorials}} | {{documentation/{{documentation/version}}/extension-section|Tutorials}} | ||
| − | * ''' | + | |
| + | === Comprehensive tutorials === | ||
| + | |||
| + | * '''Image-guided radiation therapy tutorial 2019''' ''(recommended)'' | ||
| + | ** [https://github.com/SlicerRt/SlicerRtDoc/blob/master/tutorials/SlicerRT_Tutorial_IGRT_4.11.pptx Slides including dataset] | ||
| + | * '''World Congress 2015 tutorial''' | ||
** Tutorial presentation: [https://www.dropbox.com/s/b7qx3n10s52o5f8/SlicerRT_WorldCongress_TutorialIGRT.pptx?dl=0 pptx] [https://github.com/SlicerRt/SlicerRtDoc/raw/master/tutorials/SlicerRT_WorldCongress_TutorialIGRT.pdf pdf] | ** Tutorial presentation: [https://www.dropbox.com/s/b7qx3n10s52o5f8/SlicerRT_WorldCongress_TutorialIGRT.pptx?dl=0 pptx] [https://github.com/SlicerRt/SlicerRtDoc/raw/master/tutorials/SlicerRT_WorldCongress_TutorialIGRT.pdf pdf] | ||
** Dataset: [http://slicer.kitware.com/midas3/download/item/205391/WC2015_Gel_Slicelet_Dataset.zip download] from MIDAS | ** Dataset: [http://slicer.kitware.com/midas3/download/item/205391/WC2015_Gel_Slicelet_Dataset.zip download] from MIDAS | ||
| Line 108: | Line 114: | ||
* '''RSNA 2012 tutorial''' | * '''RSNA 2012 tutorial''' | ||
** Tutorial description: [http://www.donotlink.com/bEo SlicerRT wiki: Slicer tutorials at RSNA 2012] | ** Tutorial description: [http://www.donotlink.com/bEo SlicerRT wiki: Slicer tutorials at RSNA 2012] | ||
| − | |||
** Sample data: [http://slicer.kitware.com/midas3/folder/859 download] SlicerRT ART dose verification data from Midas server | ** Sample data: [http://slicer.kitware.com/midas3/folder/859 download] SlicerRT ART dose verification data from Midas server | ||
| − | * | + | |
| − | * | + | === Module tutorials === |
| − | + | ||
| + | * [https://github.com/SlicerRt/SlicerRtDoc/blob/master/tutorials/SlicerRT_Tutorial_OrthovoltageDoseEngine.pptx External beam planning tutorial for orthovoltage RT] (uses EGSnrc) | ||
| + | * [https://github.com/SlicerRt/SlicerRtDoc/blob/master/tutorials/SlicerRT_Tutorial_DoseSurfaceHistogram.pptx Dose surface histogram tutorial] | ||
| + | * [https://github.com/SlicerRt/SlicerRtDoc/blob/master/tutorials/SlicerRT_Tutorial_Isodose.pptx Isodose tutorial] | ||
| + | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
Latest revision as of 02:26, 27 November 2019
Home < Documentation < Nightly < Extensions < SlicerRT
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
Authors: Csaba Pinter (PerkLab, Queen's University), Andras Lasso (PerkLab, Queen's University)
Contributors: Greg Sharp (Massachusetts General Hospital), Kevin Wang (Princess Margaret Hospital, UHN Toronto), Steve Pieper (Isomics)
Contacts:
- Csaba Pinter, <email>pinter.csaba@gmail.com</email>
- Andras Lasso, <email>lasso@cs.queensu.ca</email>
- Slicer Forum
- How to report an error
Website: slicerrt.org
License: Slicer license
Download/install: install 3D Slicer, start 3D Slicer, open the Extension Manager, install the SlicerRT extension (see more details on the download page)
Extension Description
|
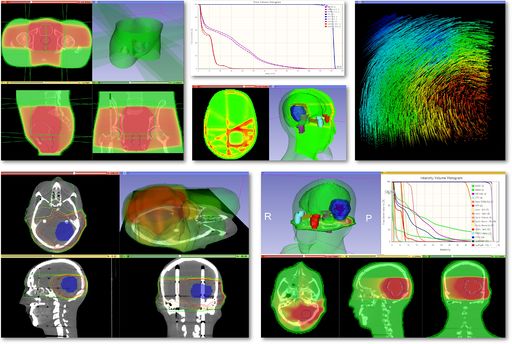
Modules
|
|
Use Cases
- Comparison of dose maps and dose volume histograms from various treatment planning systems
- Evaluation of the effect of different adaptive techniques (IGRT, image-based non-rigid patient motion compensation, etc.)
- Calculate couch shift parameters for patient setup correction in IGRT
- Dose accumulation with motion compensation
- Testing of treatment planning algorithms
- Calculation of PTV margin
- Proton dose calculation
- Gel dosimetry analysis
- Tumor volume tracking
- Treatment plan similarity measurement in the cloud
- Batch structure set conversion
Tutorials
Comprehensive tutorials
- Image-guided radiation therapy tutorial 2019 (recommended)
- World Congress 2015 tutorial
- Summer NA-MIC week 2013 tutorial
- ECR 2013 - Medical University Vienna workshop
- Workshop material: download from NA-MIC.org
- RSNA 2012 tutorial
- Tutorial description: SlicerRT wiki: Slicer tutorials at RSNA 2012
- Sample data: download SlicerRT ART dose verification data from Midas server
Module tutorials
- External beam planning tutorial for orthovoltage RT (uses EGSnrc)
- Dose surface histogram tutorial
- Isodose tutorial
Similar Extensions
- Plastimatch: SlicerRT and Plastimatch are complementary software libraries. Plastimatch focuses on delivering new computational methods radiotherapy, while SlicerRT aims for providing an easy-to-use interface for a wide range of stable, well-tested radiotherapy related features. SlicerRT uses Plastimatch internally for certain operations.
- Gel Dosimetry: Slicelet facilitating a streamlined workflow to perform true 3D gel dosimetry analysis for commissioning linacs and evaluating new dose calculation procedures
- Film Dosimetry: Slicelet supporting workflow to perform 2D film dosimetry analysis for commissioning new radiation techniques and to validate the accuracy of radiation treatment by enabling visual comparison of the planned dose to the delivered dose
References
How to cite
Please cite the following paper when referring to SlicerRt in your publication:
C. Pinter, A. Lasso, A. Wang, D. Jaffray and G. Fichtinger, "SlicerRT – Radiation therapy research toolkit for 3D Slicer", Med. Phys., 39(10) pp. 6332-6338, 2012
@ARTICLE{Pinter2012,
author = {Pinter, C. and Lasso, A. and Wang, A. and Jaffray, D. and Fichtinger, G.},
title = {SlicerRT – Radiation therapy research toolkit for 3D Slicer},
journal = {Med. Phys.},
year = {2012},
volume = {39},
number = {10},
pages = {6332-6338},
}




