Difference between revisions of "Documentation/4.6/Training"
From Slicer Wiki
Tag: 2017 source edit |
|||
| (29 intermediate revisions by 5 users not shown) | |||
| Line 1: | Line 1: | ||
| − | <noinclude>{{documentation/ | + | <noinclude>{{documentation/historicaltraining}} |
=Introduction: Slicer {{documentation/version}} Tutorials= | =Introduction: Slicer {{documentation/version}} Tutorials= | ||
| Line 145: | Line 145: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
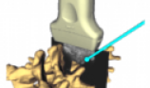
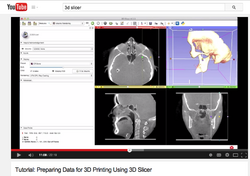
| − | * This | + | * This [https://www.youtube.com/watch?v=MKLWzD0PiIc 3D Printing Tutorial] is a video-based tutorial that shows how to prepare 3D Slicer data for 3D printing using Slicer4.3. |
| − | * | + | ** Author: Nabgha Farhat, MSc |
| − | * Audience: Users and developers interested in 3D printing | + | * The [[Documentation/{{documentation/version}}/Training#Segmentation_for_3D_printing|Segmentation for 3D printing Step-by-step tutorial]] shows how to use the Segment Editor of 3D Slicer for 3D printing using Slicer4.6. |
| + | ** Author: Csaba Pinter, MSc | ||
| + | ** Audience: Users and developers interested in 3D printing | ||
|align="right"|[[Image:3DPrinting_tutorial.png|250px]] | |align="right"|[[Image:3DPrinting_tutorial.png|250px]] | ||
|} | |} | ||
| Line 174: | Line 176: | ||
|} | |} | ||
| + | ==Slicer4 Radiation Therapy Tutorial == | ||
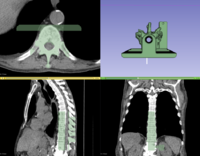
| + | ** The [https://github.com/SlicerRt/SlicerRtDoc/raw/master/tutorials/SlicerRT_WorldCongress_TutorialIGRT.pdf SlicerRT tutorial] is an introduction to the Radiation Therapy functionalities of Slicer. | ||
| + | ** Author: Csaba Pinter, Andras Lasso, An Wang, Gregory C. Sharp, David Jaffray, Gabor Fichtinger. | ||
| + | ** Dataset: [http://slicer.kitware.com/midas3/download/item/205404/SlicerRT_WorldCongress_TutorialIGRT_Dataset.zip download] from MIDAS | ||
| + | **Based on Slicer 4.6 | ||
== Other == | == Other == | ||
| Line 186: | Line 193: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *The [ | + | *The [https://github.com/SlicerRt/SlicerRtDoc/raw/master/tutorials/SegmentationFor3DPrinting_TutorialContestWinter2017.pdf Segmentation for 3D printing Tutorial] ([https://github.com/SlicerRt/SlicerRtDoc/raw/master/tutorials/SegmentationFor3DPrinting_TutorialContestWinter2017.pptx pptx]) is an introduction to the new [[Documentation/{{documentation/version}}/Modules/SegmentEditor|Segment Editor]] module of Slicer. |
*Author: Csaba Pinter (Queen's University, Canada) | *Author: Csaba Pinter (Queen's University, Canada) | ||
| − | *Dataset: | + | *Dataset: [[:File:BasePiece.zip|Phantom base STL model]] Source: [http://perk-software.cs.queensu.ca/plus/doc/nightly/modelcatalog/ PerkLab]. |
|align="right"| | |align="right"| | ||
[[File:SlicerWinterProjectWeek2017-Segmentation-for-3d-printing.png | 200px]]. | [[File:SlicerWinterProjectWeek2017-Segmentation-for-3d-printing.png | 200px]]. | ||
| Line 196: | Line 203: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
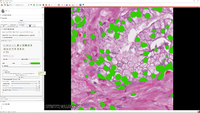
| − | *The [[Documentation/{{documentation/version}}/Extensions/SlicerPathology|Slicer Pathology Tutorial]] describes how to use the | + | *The [[Documentation/{{documentation/version}}/Extensions/SlicerPathology|Slicer Pathology Tutorial]] describes how to use the automated and semi-automated segmentation tools of the Slicer Pathology Extension. |
*Author: Erich Bremer (Stonybrook), Andriy Fedorov (Brigham and Women’s Hospital) | *Author: Erich Bremer (Stonybrook), Andriy Fedorov (Brigham and Women’s Hospital) | ||
| − | |||
|align="right"| | |align="right"| | ||
[[File:SlicerPathologyScreenShot8.png | 200px]]. | [[File:SlicerPathologyScreenShot8.png | 200px]]. | ||
| Line 206: | Line 212: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *The [https://www.na-mic.org/Wiki/index.php/File:SimpleDiffusionGradientInformationExtractorTutorial_Chauvin_Jan2017.pdf Simple Multi-shell Diffusion Gradients Information Extractor Tutorial] describes how to use a simple Python script for parsing multi-shell sensitizing gradients information from | + | *The [https://www.na-mic.org/Wiki/index.php/File:SimpleDiffusionGradientInformationExtractorTutorial_Chauvin_Jan2017.pdf Simple Multi-shell Diffusion Gradients Information Extractor Tutorial] describes how to use a simple Python script for parsing multi-shell sensitizing gradients information from diffusion MRI datasets in Nifti file. |
*Author: Laurent Chauvin (ETS Montreal) | *Author: Laurent Chauvin (ETS Montreal) | ||
|align="right"| | |align="right"| | ||
| Line 215: | Line 221: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *The [https://www.nitrc.org/docman/view.php/308/1982/SPHARM-PDM_Tutorial_July2015.pdf SPHARM-PDM Tutorial] describes how to use SPHARM-PDM and ShapePopulationViewer Slicer extensions to | + | *The [https://www.nitrc.org/docman/view.php/308/1982/SPHARM-PDM_Tutorial_July2015.pdf SPHARM-PDM Tutorial] describes how to use the SPHARM-PDM and ShapePopulationViewer Slicer extensions to compute point-based models for Shape Analysis and perform quality control of the different models. |
*Author: Jonathan Perdomo (UNC), Beatriz Paniagua (Kitware Inc.) | *Author: Jonathan Perdomo (UNC), Beatriz Paniagua (Kitware Inc.) | ||
*Dataset: [https://www.nitrc.org/docman/view.php/308/1981/SPHARM_Tutorial_Data_July2015.zip Tutorial Data] | *Dataset: [https://www.nitrc.org/docman/view.php/308/1981/SPHARM_Tutorial_Data_July2015.zip Tutorial Data] | ||
| Line 225: | Line 231: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *The [https://www.na-mic.org/Wiki/index.php/File:ROSIGTLTutorial. | + | *The [https://www.na-mic.org/Wiki/index.php/File:ROSIGTLTutorial.pdf Integration of Robot Operating System (ROS) and 3D Slicer using OpenIGTLink Tutorial] is an introduction to the software architecture of surgical robot systems. The tutorial provides hands-on experience on the integration of software and hardware for medical robotics applications. |
*Author: Junichi Tokuda (Brigham and Women’s Hospital) | *Author: Junichi Tokuda (Brigham and Women’s Hospital) | ||
*Dataset: Not available. | *Dataset: Not available. | ||
| Line 235: | Line 241: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
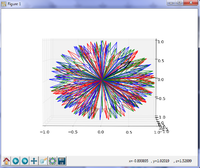
| − | *The [ | + | *The [https://www.na-mic.org/Wiki/index.php/File:Fiber_Bundle_Volume_Measurement_TutorialContestWinter2017.pdf Fiber Bundle Volume Measurement Tutorial] shows how to calculate the volume of tractography reconstructions of white matter tracts. |
*Author: Shun Gong (Shanghai Changzheng Hospital, China) | *Author: Shun Gong (Shanghai Changzheng Hospital, China) | ||
| − | *Dataset: [http://www.na-mic.org/Wiki/images/4/4c/FiberVolume_data.zip Tutorial data] | + | *Dataset: [http://www.na-mic.org/Wiki/images/4/4c/FiberVolume_data.zip Tutorial data] |
|align="right"| | |align="right"| | ||
[[File:SlicerWinterProjectWeek2017-FiberBundleVolumeMeasurements.png | 200px]]. | [[File:SlicerWinterProjectWeek2017-FiberBundleVolumeMeasurements.png | 200px]]. | ||
|} | |} | ||
| − | |||
==Winter 2016 Tutorial contest== | ==Winter 2016 Tutorial contest== | ||
| Line 391: | Line 396: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *[http:// | + | *[http://www.slicer.org/w/img_auth.php/f/f1/OpenIGTLinkTutorial_Slicer4.1.0_JunichiTokuda_Apr2012.pdf OpenIGTLink] |
*Authors: Junichi Tokuda, BWH | *Authors: Junichi Tokuda, BWH | ||
|align="right"| | |align="right"| | ||
| Line 425: | Line 430: | ||
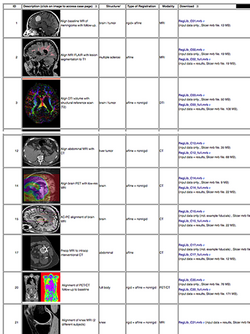
:Author: Dominik Meier, Ph.D. | :Author: Dominik Meier, Ph.D. | ||
:Audience: users interested learning/applying Slicer image registration technology | :Audience: users interested learning/applying Slicer image registration technology | ||
| − | |align="right"|[[Image:RegLib_table.png|250px|link= | + | |align="right"|[[Image:RegLib_table.png|250px|link=https://www.slicer.org/wiki/Documentation/{{documentation/version}}/Registration/RegistrationLibrary]]|} |
| − | |} | ||
---- | ---- | ||
<br> | <br> | ||
| Line 439: | Line 443: | ||
= External Resources = | = External Resources = | ||
| − | == Using the Editor == | + | == Murat Maga's blog posts about using 3D Slicer for biology == |
| + | |||
| + | * [https://blogs.uw.edu/maga/2017/04/11/getting-started-with-3d-slicer-as-a-biologist/ Slicer for Biologists] | ||
| + | * [https://blogs.uw.edu/maga/2017/04/11/a-worked-example-getting-and-visualizing-data-from-digimorph/ Loading data from DigiMorph] | ||
| + | * [https://blogs.uw.edu/maga/2017/04/11/morphosource-data-and-dealing-with-dicom-series-in-slicer/ Fixing problem DICOM] | ||
| + | * [https://blogs.uw.edu/maga/2017/04/12/scissors-tool-is-awesome/ Scissors tool is awesome] | ||
| + | |||
| + | == Using the (legacy) Editor == | ||
This set of tutorials about the use of slicer in paleontology is very well written and provides step-by-step instructions. Even though it covers slicer version 3.4, many of the concepts and techniques have applicability to the new version and to any 3D imaging field: | This set of tutorials about the use of slicer in paleontology is very well written and provides step-by-step instructions. Even though it covers slicer version 3.4, many of the concepts and techniques have applicability to the new version and to any 3D imaging field: | ||
Latest revision as of 22:12, 22 November 2022
Home < Documentation < 4.6 < Training
 |
This section is currently out-of-date and may contain errors but is retained for historical reference. Up-to-date training materials can be found at Documentation/Nightly/Training |
Introduction: Slicer 4.6 Tutorials
- The 3D Slicer compendium is a collection of hands-on tutorials with anonymized sample data sets. The tutorials demonstrate how to use the 3D Slicer software platform (version 4.6 release) to accomplish certain tasks.
- For tutorials for other versions of Slicer, please visit the Slicer training portal.
- For "reference manual" style documentation, please visit the Slicer 4.6 documentation page
- Some of these tutorials are based on older releases of 3D Slicer. The concepts are still useful but bear in mind that some interface elements and features will be different in updated versions.
- For general questions related to the Slicer4 Training Compendium and to 3D Slicer training events, please send an e-mail to Sonia Pujol, Ph.D, Director of Training of 3D Slicer
Contents
- 1 Introduction: Slicer 4.6 Tutorials
- 2 General Introduction
- 3 Tutorials for Software Developers
- 4 Specific functions
- 4.1 Slicer4 Diffusion Tensor Imaging Tutorial
- 4.2 Slicer4 Neurosurgical Planning Tutorial
- 4.3 Slicer4 3D Visualization of DICOM Images for Radiology Applications
- 4.4 Slicer4 Quantitative Imaging Tutorial
- 4.5 Slicer4 IGT
- 4.6 Slicer4 3D Printing
- 4.7 Slicer4 Image Registration
- 4.8 Fast GrowCut
- 4.9 Slicer4 Radiation Therapy Tutorial
- 4.10 Other
- 5 3D Slicer Tutorial contests
- 6 Additional resources
- 7 External Resources
General Introduction
Slicer Welcome Tutorial
|
Slicer4Minute Tutorial
|
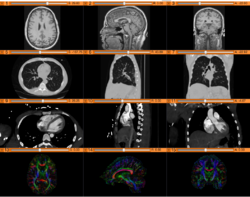
Slicer4 Data Loading and 3D Visualization
|
Tutorials for Software Developers
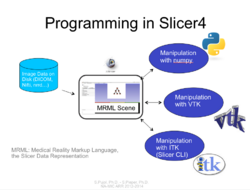
Slicer4 Programming Tutorial
|
For additional Python scripts examples, please visit the Script Repository page
Developing and Contributing Extensions for 3D Slicer
|
Specific functions
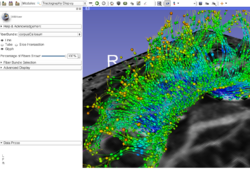
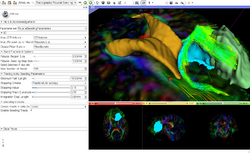
Slicer4 Diffusion Tensor Imaging Tutorial
|
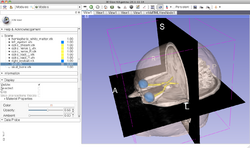
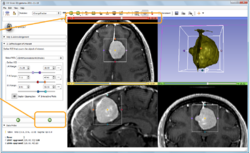
Slicer4 Neurosurgical Planning Tutorial
|
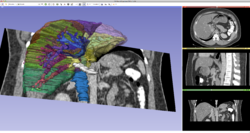
Slicer4 3D Visualization of DICOM Images for Radiology Applications
|
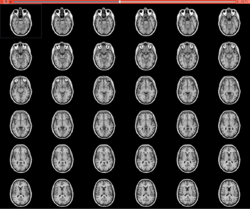
Slicer4 Quantitative Imaging Tutorial
|
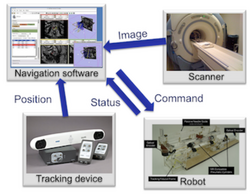
Slicer4 IGT
|
Slicer4 3D Printing
|
|
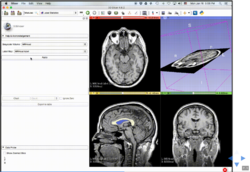
Slicer4 Image Registration
|
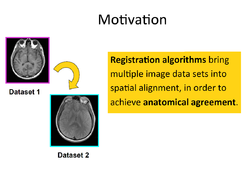
|
- Based on: 3D Slicer version 4.5
See the Registration Library for worked out registration examples with data.
Fast GrowCut
|

|
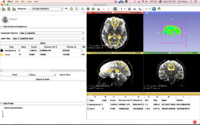
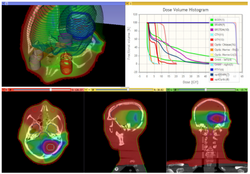
Slicer4 Radiation Therapy Tutorial
- The SlicerRT tutorial is an introduction to the Radiation Therapy functionalities of Slicer.
- Author: Csaba Pinter, Andras Lasso, An Wang, Gregory C. Sharp, David Jaffray, Gabor Fichtinger.
- Dataset: download from MIDAS
- Based on Slicer 4.6
Other
Additional (non-curated) videos-based demonstrations using 3D Slicer are accessible on You Tube.
3D Slicer Tutorial contests
Winter 2017 Tutorial contest
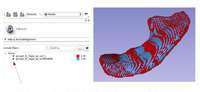
Segmentation for 3D printing
|
Slicer Pathology
|
Simple Python Tool for Quality Control of DWI data
|
SPHARM-PDM
|
Integration of Robot Operating System (ROS) and 3D Slicer using OpenIGTLink
|
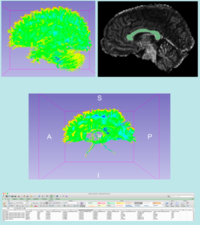
Fiber Bundle Volume Measurement
|
Winter 2016 Tutorial contest
Subject Hierarchy
|
Fiber Bundle Selection and Scalar Measurements
|
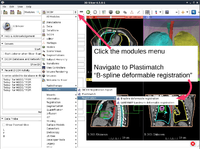
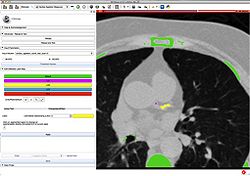
Plastimatch
|
UKF
|
Summer 2014 Tutorial contest
Cardiac Agatston Tutorial
|
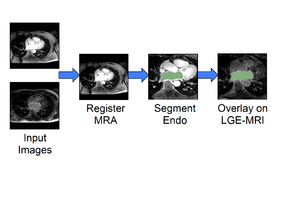
CMR Toolkit LA workflow
|
Summer 2013 Tutorial contest
Cardiac MRI Toolkit
|
HelloCLI
|
SlicerRT
|
DTIPrep
|
Summer 2012 Tutorial contest
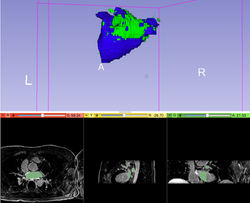
Automatic Left Atrial Scar Segmenter
|
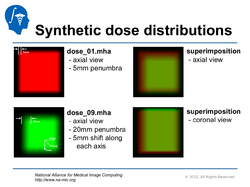
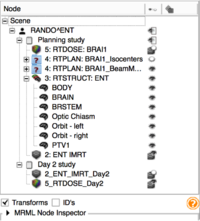
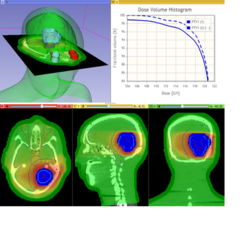
Qualitative and Quantitative Comparison of Two RT Dose Distributions
|
Dose Accumulation for Adaptive Radiation Therapy
|
WebGL Export
|
OpenIGTLink
|
Additional resources
|
|
|
|
|
 |} |}
External ResourcesMurat Maga's blog posts about using 3D Slicer for biologyUsing the (legacy) EditorThis set of tutorials about the use of slicer in paleontology is very well written and provides step-by-step instructions. Even though it covers slicer version 3.4, many of the concepts and techniques have applicability to the new version and to any 3D imaging field:
Team ContributionsSee the collection of videos on the Kitware vimeo album. User ContributionsSee the User Contributions Page for more content. |