Difference between revisions of "EMSegmenter-Tasks:MRI-Human-Brain-Parcellation"
| Line 53: | Line 53: | ||
For each case from previous manually segmented 1.5T scans (Nakamura et al, 2007), binary label maps for the 20 leaves (BG,ltgm1-ltgm4, rtgm1-rtgm4, subgm, ltwm1-ltwm4, rtwm1-rtwm4, subwm and CSF) to be segmented were obtained. Initially, all these label maps were respectively affine transformed to a pre-determined case. A diffeomorphic registration algorithm (Vercauteren et al, 2008) was applied and this produced warps of all the regions for all the cases. The warps were applied to the original images and all the images for respective regions are aligned in a final atlas space. Probability maps are then determined by averaging the images for respective regions i.e. BG, ltgm1, ltgm2, CSF etc. | For each case from previous manually segmented 1.5T scans (Nakamura et al, 2007), binary label maps for the 20 leaves (BG,ltgm1-ltgm4, rtgm1-rtgm4, subgm, ltwm1-ltwm4, rtwm1-rtwm4, subwm and CSF) to be segmented were obtained. Initially, all these label maps were respectively affine transformed to a pre-determined case. A diffeomorphic registration algorithm (Vercauteren et al, 2008) was applied and this produced warps of all the regions for all the cases. The warps were applied to the original images and all the images for respective regions are aligned in a final atlas space. Probability maps are then determined by averaging the images for respective regions i.e. BG, ltgm1, ltgm2, CSF etc. | ||
| + | Weighted combined atlases: Left and right classification of the brain typically relies on the atlases since the intensities are similar. Thus if an area on the left side say ltgm1 is not covered adequately by the atlas, the segmentation algorithm puts the next region with the highest probability in the ltgm1 region. Therefore, in order to get an accurate segmentation and to prevent left and right mis-classification, the atlases were weighted and combined for each leaf. For example, for ltgm1 the ltgm1 atlas was weighted by 0.9 and the ltwm1 atlas was weighted by 0.1 and then summed up. Thus in this way, for ltgm1 the atlas covers even the white matter region except that the probability of finding grey matter is lower. The opposite if done for ltwm1 i.e. the ltwm1 atlas is weighted by 0.9 and ltgm1 is weighted by 0.1 and then summed up. These weighted combined atlases provide a much accurate segmentation. | ||
Revision as of 17:49, 9 December 2010
Home < EMSegmenter-Tasks:MRI-Human-Brain-ParcellationReturn to EMSegmenter Task Overview Page
Contents
Description
MRI Human Head pipeline for a finer-grained parcellation (TODO: Priya please provide description and proper citation)
The pipeline consist of the following steps:
- Step 1: Perform image inhomogeneity correction of the MRI scan via N4ITKBiasFieldCorrection (Tustison et al 2010)
- Step 2: Register the atlas to the MRI scan via BRAINSFit (Johnson et al 2007)
- Step 3: Compute the intensity distributions for each structure
Compute intensity distribution (mean and variance) for each label by automatically sampling from the MR scan. The sampling for a specific label is constrained to the region that consists of voxels with high probability (top 95%) of being assigned to the label according to the aligned atlas.
- Step 4: Automatically segment the MRI scan into the structures of interest using EM Algorithm (Pohl et al 2007)
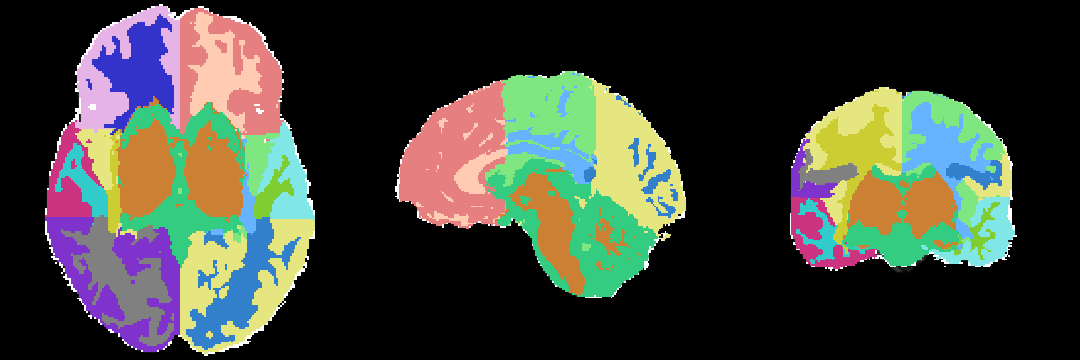
Anatomical Tree
The anatomical tree represents the structures to be segmented. Node labels displayed below contain a human readable structure name and in parentheses the internally used structure name.
- Root
- background (BG)
- grey matter (GM)
- left grey matter (LTGM)
- left grey matter - region 1 (LTGM1)
- left grey matter - region 2 (LTGM2)
- left grey matter - region 3 (LTGM3)
- left grey matter - region 4 (LTGM4)
- right grey matter (RTGM)
- right grey matter - region 1 (RTGM1)
- right grey matter - region 2 (RTGM2)
- right grey matter - region 3 (RTGM3)
- right grey matter - region 4 (RTGM4)
- subcortical grey matter (SUBGM)
- left grey matter (LTGM)
- white matter (WM)
- left white matter (LTWM)
- left white matter - region 1 (LTWM1)
- left white matter - region 2 (LTWM2)
- left white matter - region 3 (LTWM3)
- left white matter - region 4 (LTWM4)
- right white matter (RTWM)
- right white matter - region 1 (RTWM1)
- right white matter - region 2 (RTWM2)
- right white matter - region 3 (RTWM3)
- right white matter - region 4 (RTWM4)
- subcortical white matter (SUBWM)
- left white matter (LTWM)
- cerebrospinal fluid (CSF)
Atlas
The atlas has been created by Padmapriya Srinivasan and Ryan Eckbo and Sylvain Bouix from (PNL-BWH)
Image Dimension = 256 x 256 x 220
Image Spacing = 0.9375 x 0.9375 x 0.9375
Briefly, the steps followed for creating the atlases are described here:
For each case from previous manually segmented 1.5T scans (Nakamura et al, 2007), binary label maps for the 20 leaves (BG,ltgm1-ltgm4, rtgm1-rtgm4, subgm, ltwm1-ltwm4, rtwm1-rtwm4, subwm and CSF) to be segmented were obtained. Initially, all these label maps were respectively affine transformed to a pre-determined case. A diffeomorphic registration algorithm (Vercauteren et al, 2008) was applied and this produced warps of all the regions for all the cases. The warps were applied to the original images and all the images for respective regions are aligned in a final atlas space. Probability maps are then determined by averaging the images for respective regions i.e. BG, ltgm1, ltgm2, CSF etc.
Weighted combined atlases: Left and right classification of the brain typically relies on the atlases since the intensities are similar. Thus if an area on the left side say ltgm1 is not covered adequately by the atlas, the segmentation algorithm puts the next region with the highest probability in the ltgm1 region. Therefore, in order to get an accurate segmentation and to prevent left and right mis-classification, the atlases were weighted and combined for each leaf. For example, for ltgm1 the ltgm1 atlas was weighted by 0.9 and the ltwm1 atlas was weighted by 0.1 and then summed up. Thus in this way, for ltgm1 the atlas covers even the white matter region except that the probability of finding grey matter is lower. The opposite if done for ltwm1 i.e. the ltwm1 atlas is weighted by 0.9 and ltgm1 is weighted by 0.1 and then summed up. These weighted combined atlases provide a much accurate segmentation.
|
|
|
|
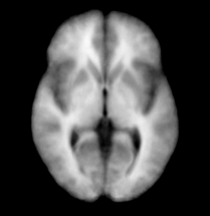
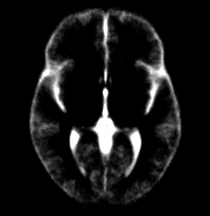
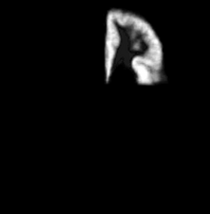
| Template (T1) | CSF | Left GM | Left WM |
Result
Collaborators
Padmapriya Srinivasan and Sylvain Bouix (PNL-BWH)
Acknowledgment
The construction of the pipeline was supported by funding from NIH NCRR 2P41RR013218 Supplement.
Citations
- Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, Gee JC N4ITK: Improved N3 Bias Correction, IEEE Trans Med Imag, 2010
- Pohl K, Bouix S, Nakamura M, Rohlfing T, McCarley R, Kikinis R, Grimson W, Shenton M, Wells W. A Hierarchical Algorithm for MR Brain Image Parcellation. IEEE Transactions on Medical Imaging. 2007 Sept;26(9):1201-1212.
- S. Warfield, J. Rexilius, P. Huppi, T. Inder, E. Miller, W. Wells, G. Zientara, F. Jolesz, and R. Kikinis, “A binary entropy measure to assess nonrigid registration algorithms,” in MICCAI, LNCS, pp. 266–274, Springer, October 2001.
- Johnson H.J., Harris G., Williams K. BRAINSFit: Mutual Information Registrations of Whole-Brain 3D Images, Using the Insight Toolkit, The Insight Journal, July 2007
- M. Nakamura, D. F. Salisbury, Y. Hirayasu, S. Bouix, K. Pohl, T. Yoshida, M. Koo, M. Koo, R. McCarley. Neocortical Gray Matter Volume in First-Episode Schizophrenia and First-Episode Affective Psychosis: A Cross-Sectional and Longitudinal MRI Study. Biological Psychiatry .Volume 62, Number 7, Pages 773-783. 2007
- T. Vercauteren, X. Pennec, A. Perchant, N. Ayache. Symmetric Log-Domain Diffeomorphic Registration: A Demons-based Approach. MICCAI 2008